|
SARS-CoV-2 nucleic acid and protein sequences did become available shortly after the start of COVID-19 pandemic and biological aspects of these genes/proteins including their interactions with immune system have been under intense investigation. 
Viral vaccines including polio and MMR have been suggested as candidates that might give rise to this potential ‘cross-protective’ phenomenon, raising the possibility for the existence of similar molecular structures. 
Indeed, several reports have suggested that immunization against other human pathogens might influence susceptibility to SARS-CoV-2. The results of these types of analyses can go beyond ordinary phylogenetics and might have applied clinicopathological implications. One approach towards better understanding of a new pathogen is comparative analysis of its nucleic acids, proteins and molecular structures. This necessitates investigations that can provide a better insight regarding coronavirus genes, proteins and biological behavior. Nonetheless, emergence of three deadly coronavirus epidemics over the last 20 years has challenged this view and is likely indicative of the dynamic evolution of these pathogens. In the first few decades after their discovery, and when compared with other major viral pathogens like smallpox, polio and influenza, coronaviruses seemed to display a more benign attitude towards their hosts. First human coronaviruses were discovered in 1960s, but human coronavirus infections have likely existed for centuries. These data indicate that there might be a degree of convergent evolution between SARS-CoV-2 and paramyxovirus surface proteins which could be of pathogenic and immunological importance.Ĭurrent COVID-19 pandemic which is caused by a newly emerged betacoronavirus has led to immense health and socio-economic problems around the world. Epitope analysis using experimentally verified data deposited in Immune Epitope Database (IEDB) revealed that several B cell epitopes as well as T cell and MHC binding epitopes map within the spike-fusion similarity region. The ‘spike-fusion’ similarity was also observed for some pathogenic coronaviruses other than SARS-CoV-2. We next analyzed spike proteins from SARS-CoV-2 “variants of concern” (VOCs) and “variants of interests” (VOIs) and found that some of these variants show considerably higher spike-fusion similarity with paramyxoviruses. This similarity was restricted to a segment of spike protein S2 subunit which is involved in cell fusion. spike) protein with paramyxovirus fusion proteins. We also observed similarities between SARS-CoV-2 surface (i.e. Comparing the proteins of SARS-CoV-2 with non-coronavirus positive and negative strand ssRNA viruses revealed multiple sequence similarities between SARS-CoV-2 and non-coronaviruses, including similarities between RNA-dependent RNA-polymerases and helicases (two highly-conserved proteins). 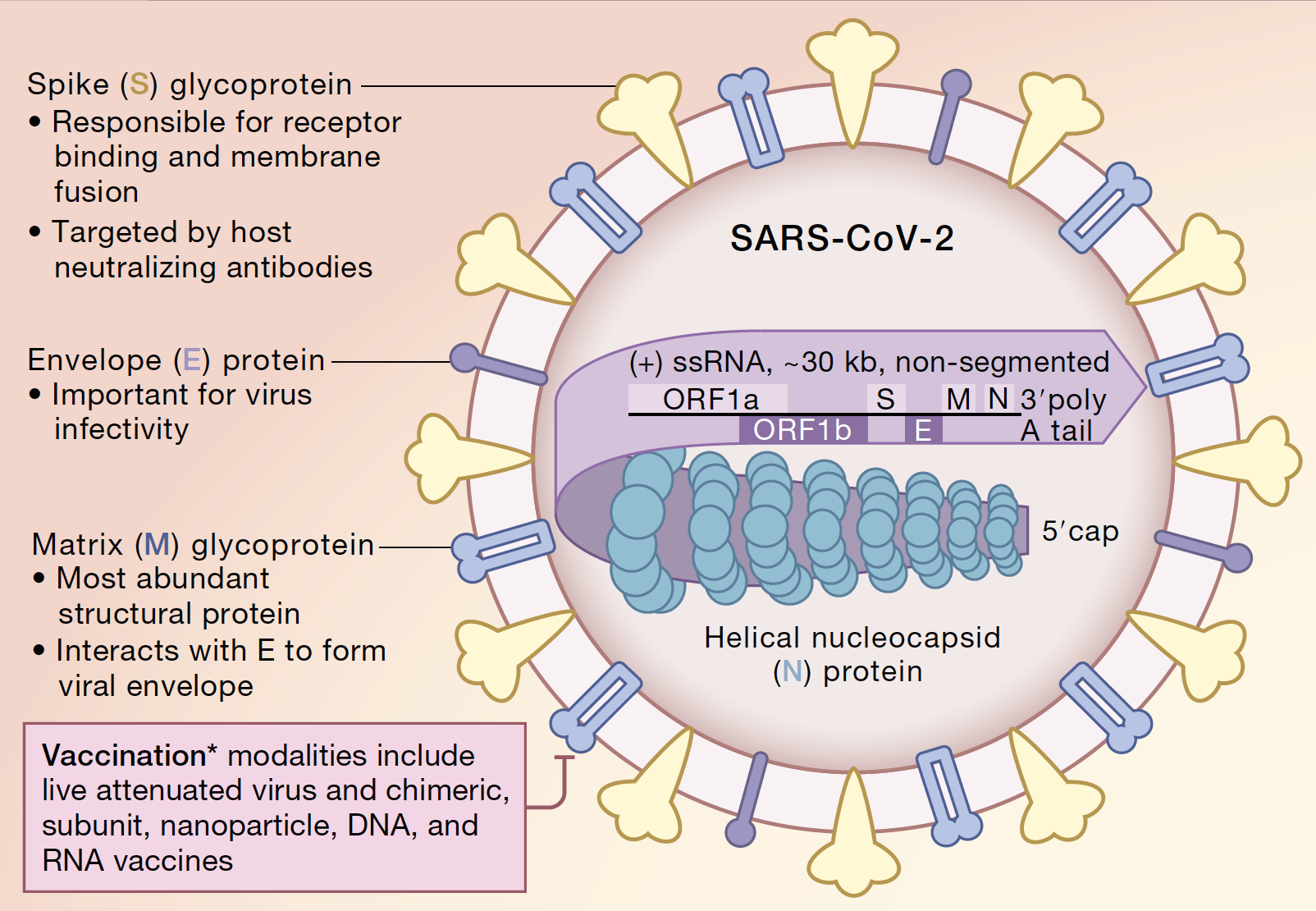
In this study, we performed bioinformatic analyses on SARS-CoV-2 protein sequences, trying to unravel potential molecular similarities between this newly emerged pathogen with non-coronavirus ssRNA viruses. Recent emergence of SARS-CoV-2 and associated COVID-19 pandemic have posed a great challenge for the scientific community.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
